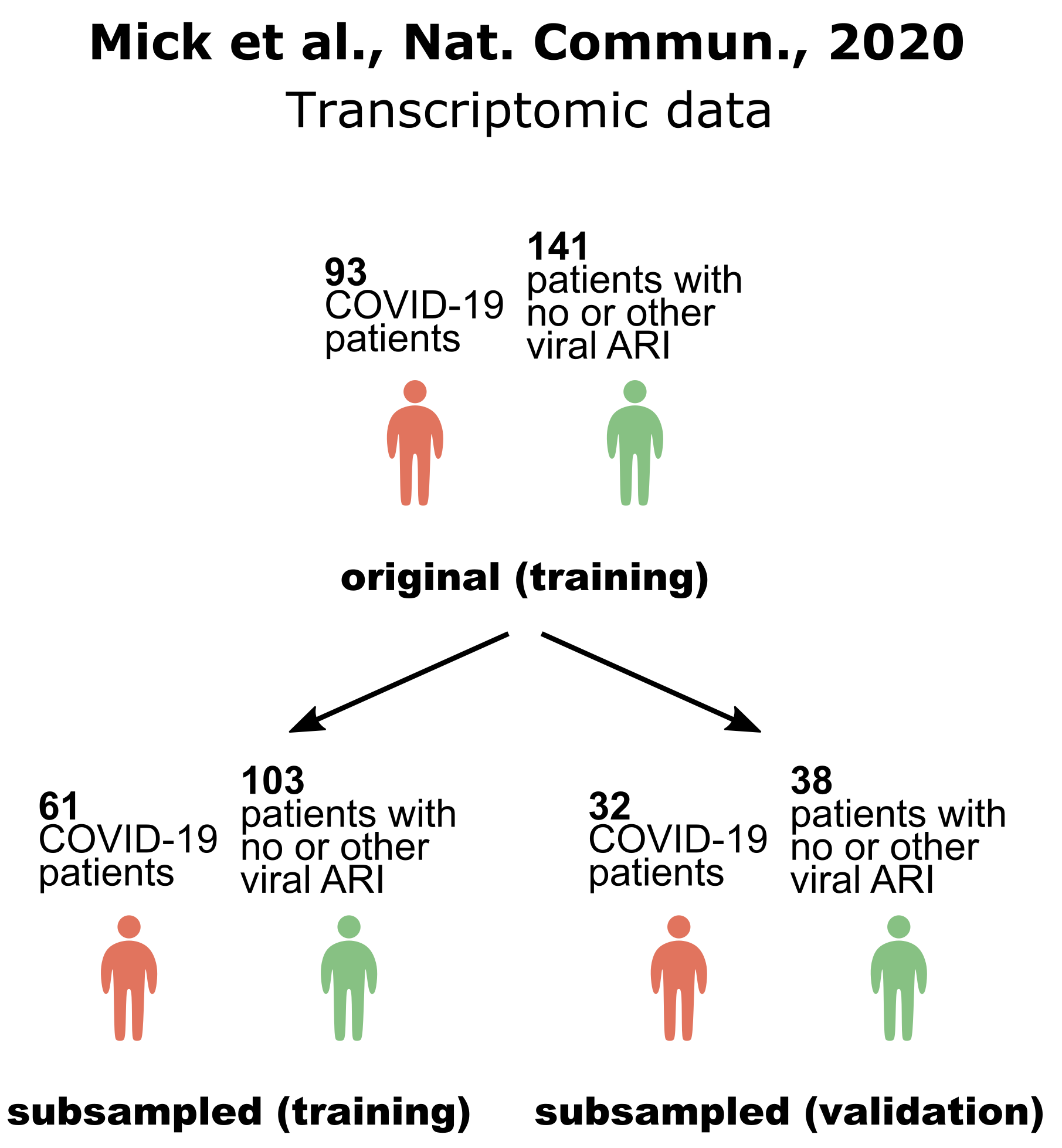
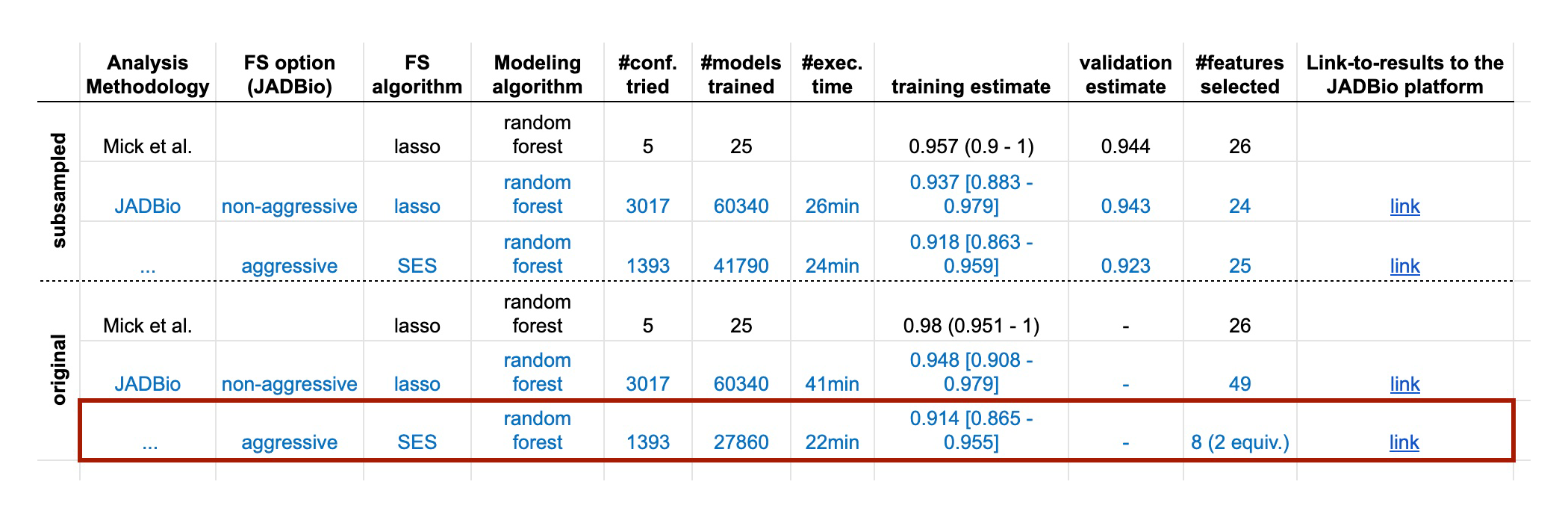
Fig. 2 Predictive performance estimates in terms of AUC reported in the original dataset publication, by JADBio on the full set of available data (i.e. no samples are lost to estimation) and by JADBio on training and validation sets (subsampled). Numbers in parentheses denote the range of the estimate while the numbers in brackets the 95% confidence intervals. The equivalences denote the number of equivalent signatures found by JADBio, e.g., “8 (2 equiv.)” means that JADBio discovered 2 equivalent signatures each containing 8 biomarkers. Each link to the JADBio platform leads to a report with the complete list of AutoML results. JADBio does not overestimate when there are no samples held out for estimation; confirms the predicted performance obtained in the original publication and discovers novel signatures.
DISCUSSION
Can AutoML improve the diagnostic/predictive models for COVID-19?
In this study, we applied AutoML in order to obtain accurate diagnostic/predictive models for COVID-19, using available archived datasets. We asked; could we improve on the predictive power of the models? Can we reduce the number of measurements required without sacrificing performance to develop a cost-effective laboratory test? Can we obtain more accurate training estimates that better reflect the performance anticipated in a real life setting? Most importantly, can AutoML improve on these aspects in a fully automated mode?
Using autoML, we have affirmatively answered all these research questions. That is, our approach was on par or better than the published results or the ones obtained by running previously used code and methodology on our training sets. Quite importantly, the respective predictive performance estimates accurately reflect the performance obtained on the validation sets, so we argue that there is no need to lose samples to estimation. JADBio internally handles estimation techniques, so that the user does not have to worry about this: it performs cross-validation, repeats the cross-validation with different fold partitions for low sample size to reduce variance of estimation, stratifies the partitioning to folds of cross-validation to reduce the variance of estimation and handle imbalanced data, corrects performance estimate for the “winner’s curse” and trying multiple algorithms using the bootstrap bias correction for CV, and includes all steps of the analysis (e.g., feature selection) within the cross-validation that leads to overestimation.
Thus, we advocate the use of all data for training with JADBio. Of course, this claim comes with an important disclaimer: JADBio’s theoretical guarantees of out-of-sample performance estimate hold only when the model is applied on the same data distribution. If the models are applied in a clinical setting where measurements have batch effects, the population characteristics are different, or there are other systematic differences in the data, an external validation set from that operational environment is clearly required. An additional advantage of this AutoML approach is that it is able to work in two modes; with and without aggressive feature selection. The latter may give away some predictive power to produce models with multiple (in case of biological redundancy) equivalent biosignatures of selective predictive features providing choices to the designers of diagnostic assays. Accordingly, we were able to deliver several highly diagnostic/prognostic biosignatures of minimal feature size from different types of COVID-19 data.
REFERENCES
Mick, E., Kamm, J., Pisco, A.O. et al. Upper airway gene expression reveals suppressed immune responses to SARS-CoV-2 compared with other respiratory viruses. Nat Commun 11, 5854 (2020). https://doi.org/10.1038/s41467-020-19587-y
#COVID-19 #autoML #MachineLearning, #SARS-CoV-2, #predictivemodels