This website uses cookies to improve your experience. We'll assume you're ok with this, but you can opt-out if you wish. Read More
Classification of non-TCGA cancer samples to TCGA molecular subtypes using compact feature sets
Kyle Ellrott, Christopher K. Wong, Christina Yau, Mauro A. A. Castro, Jordan A. Lee, Brian J. Karlberg, Jasleen K. Grewal, Vincenzo Lagani, Bahar Tercan, Verena Friedl, Toshinori Hinoue, Vladislav Uzunangelov, Lindsay Westlake, Xavier Loinaz, Ina Felau, Peggy I. Wang, Anab Kemal, Samantha J. Caesar-Johnson, Ilya Shmulevich, Alexander J. Lazar, Ioannis Tsamardinos, Katherine A. Hoadley, The Cancer Genome Atlas Analysis Network, A. Gordon Robertson, Theo A. Knijnenburg, Christopher C. Benz, Joshua M. Stuart, Jean C. Zenklusen, Andrew D. Cherniack, Peter W. Laird,
Joint work with the Tumor Molecular Pathology (TMP) Analysis Working Group (AWG) of the US National Institute of Health (NIH) Center for Cancer Genomics (CCG)
Digital Library: https://doi.org/10.1016/j.ccell.2024.12.002
Summary
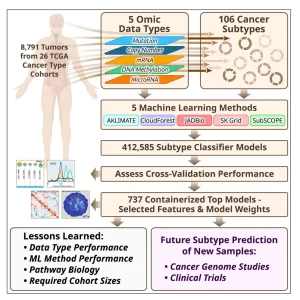
Molecular subtypes, such as defined by The Cancer Genome Atlas (TCGA), delineate a cancer’s underlying biology, bringing hope to inform a patient’s prognosis and treatment plan. However, most approaches used in the discovery of subtypes are not suitable for assigning subtype labels to new cancer specimens from other studies or clinical trials. Here, we address this barrier by applying five different machine learning approaches to multi-omic data from 8,791 TCGA tumor samples comprising 106 subtypes from 26 different cancer cohorts to build models based upon small numbers of features that can classify new samples into previously defined TCGA molecular subtypes—a step toward molecular subtype application in the clinic. We validate select classifiers using external datasets. Predictive performance and classifier-selected features yield insight into the different machine-learning approaches and genomic data platforms. For each cancer and data type we provide containerized versions of the top-performing models as a public resource.
Methodology
JADBio along with four other machine learning methods were used to train multi-omics classifiers on TCGA tumor samples. Data comprised of 5 different types: mRNA, MicroRNA, CNV, Mutation, and Methylation. The goal was to build models with as few biomarkers as possible for creating compact cancer testing panels and kits to clinically subtype non-TCGA patient tumor samples.
Results
Robust Cross-Platform Subtype Classification. The developed machine-learning models accurately classify non-TCGA cancer samples into TCGA-defined molecular subtypes across diverse tumor types and data platforms.
Minimal Feature Sets. Each classifier relies on a small, optimized set of omic features—genes, methylation sites, etc.—demonstrating that high accuracy doesn’t require large datasets, making practical panel construction feasible. Specifically, between 70 to 150 samples are required for cancer subtype classification.
High Predictive Performance. Models achieved strong precision, recall, and F1‑scores for most subtypes tested. While some very rare subtypes with minimal training samples were excluded, performance remained robust for the majority.
Resource for Panel Development. The resulting feature sets are explicitly proposed as a foundation for designing compact clinical panels or kits aimed at classifying tumor subtypes—bridging the gap between genomic profiling and personalized oncology.
Conclusions
Small sets of carefully selected multi-omic features can accurately predict cancer subtypes defined by TCGA. These compact models work well across different datasets and platforms, making them a practical public resource for use in clinical settings. This approach can help create efficient diagnostic panels that support personalized cancer treatment by linking molecular subtype information to therapeutic decisions.
How did JADBio perform?
JADBio showed strong predictive performance in 96% of cases (25/26)
- Had the highest performance in 16 out of 26 (62%) of cancer types
- Had tied best performance in 1 cancer type (Prostate, PRAD)
- Performed within 1 standard deviation from the best in 30% (8/26) of the remaining ones.
JADBio’s feature selection algorithms select the fewest biomarkers in 85% of cases (22/26)
- Rapid panel creation, no specialized expertise required.
- Unique and unbiased identification of multiple equally predictive subsets with high biological relevance.
- Non-redundant biomarker selection that improves classification using streamlined, more parsimonious biomarker sets, surpassing traditional feature ranking methods
Methods that provided the best predictive model in each of the 26 cancer types from the TCGA cohorts
Read more at Cancer Cell: Classification of non-TCGA cancer samples to TCGA molecular subtypes using compact feature sets
OTHER
Do you have questions?
JADBio can meet your needs. Ask one of our experts for an interactive demo.
Stay connected to get our news first!
Do you have questions?
JADBio can meet your needs. Ask one of our experts for an interactive demo.

Join the JADai Community!
Sign up with a FREE Basic plan! Be part of a growing community of AutoML enthusiasts
GET STARTED




